Saving Datasets and DataArrays to NetCDF¶
Objectives¶
Introduce an easy method for saving Datasets and DataArrays objects to NetCDF
Introduction¶
Saving your Datasets and DataArrays objects to NetCDF files couldn’t be simpler. The xarray module that we’ve been using to load NetCDF files provides methods for saving your Datasets and DataArrays as NetCDF files.
Here is the manual page on the subjet: http://xarray.pydata.org/en/stable/generated/xarray.Dataset.to_netcdf.html
The method ._to_netcdf( ) is available to both Datasets and DataArrays objects. So useful!
To complete this tutorial, you will need the monthly mean SSH and temperature/salinity output for March 2010 downloaded, as well as the model grid parameters file. If you have completed previous tutorials, you may have some or all of these already. The ShortNames of the required datasets are:
ECCO_L4_SSH_LLC0090GRID_MONTHLY_V4R4
ECCO_L4_TEMP_SALINITY_LLC0090GRID_MONTHLY_V4R4
ECCO_L4_GEOMETRY_LLC0090GRID_V4R4
Syntax¶
your_dataset.to_netcdf('/your_filepath/your_netcdf_filename.nc')
Saving an existing Dataset to NetCDF¶
First, let’s set up the environment and load a Dataset
[1]:
import numpy as np
import xarray as xr
import sys
import matplotlib.pyplot as plt
import json
import glob
[2]:
## Import the ecco_v4_py library into Python
## =========================================
#import ecco_v4_py as ecco
## If ecco_v4_py is not installed in your local Python library,
## tell Python where to find it. The example below adds
## ecco_v4_py to the user's path if it is stored in the folder
## ECCOv4-py under the user's home directory
from os.path import join,expanduser
user_home_dir = expanduser('~')
sys.path.append(join(user_home_dir,'ECCOv4-py'))
import ecco_v4_py as ecco
Now load a single tile (model tile 2) of monthly averaged THETA for March 2010.
[3]:
## Set top-level file directory for the ECCO NetCDF files
## =================================================================
## currently set to ~/Downloads/ECCO_V4r4_PODAAC,
## the default if ecco_podaac_download was used to download dataset granules
ECCO_dir = join(user_home_dir,'Downloads','ECCO_V4r4_PODAAC')
[4]:
# load Mar 2010 temperature/salinity monthly mean
ds_temp_sal_201003 = xr.open_dataset(glob.glob(join(ECCO_dir,'*TEMP_SALINITY*MONTHLY*','*_2010-03_*.nc'))[0])
# select only potential temperature on tile 2 and load it into memory
# Note: the extra set of brackets around THETA creates an xarray Dataset
# (with a single data variable) rather than a DataArray
theta_dataset = ds_temp_sal_201003[['THETA']].isel(tile=2).load()
[5]:
type(theta_dataset)
[5]:
xarray.core.dataset.Dataset
Now that we’ve loaded theta_dataset, let’s save it in the current file directory with a new name.
[6]:
new_filename_1 = './test_output.nc'
print ('saving to ', new_filename_1)
theta_dataset.to_netcdf(path=new_filename_1)
print ('finished saving')
saving to ./test_output.nc
finished saving
It’s really that simple!
Saving a new custom Dataset to NetCDF¶
Now let’s create a new custom Dataset that with THETA, SSH and model grid parameter variables for a few tiles.
[7]:
ds_SSH_201003 = xr.open_dataset(glob.glob(join(ECCO_dir,'*SSH*MONTHLY*','*_2010-03_*.nc'))[0])
# merge SSH and THETA data variables into a new dataset, and select several tiles to load into memory
tiles_to_load = [0,1,2]
ds_SSH_THETA_201003 = xr.merge([ds_SSH_201003['SSH'],ds_temp_sal_201003['THETA']]).isel(tile=tiles_to_load).load()
# load grid parameters (for tiles 0,1,2 only)
ds_grid = xr.open_dataset(glob.glob(join(ECCO_dir,'*GEOMETRY*','*.nc'))[0]).isel(tile=tiles_to_load).load()
custom_dataset = xr.merge([ds_SSH_THETA_201003, ds_grid])
and now we can easily save it:
[8]:
new_filename_2 = './test_output_2.nc'
print ('saving to ', new_filename_2)
custom_dataset.to_netcdf(path=new_filename_2)
custom_dataset.close()
print ('finished saving')
saving to ./test_output_2.nc
finished saving
[9]:
custom_dataset
[9]:
<xarray.Dataset>
Dimensions: (i: 90, j: 90, tile: 3, time: 1, k: 50, i_g: 90, j_g: 90, k_u: 50, k_l: 50, k_p1: 51, nb: 4, nv: 2)
Coordinates: (12/21)
* i (i) int32 0 1 2 3 4 5 6 7 8 9 10 ... 80 81 82 83 84 85 86 87 88 89
* j (j) int32 0 1 2 3 4 5 6 7 8 9 10 ... 80 81 82 83 84 85 86 87 88 89
* tile (tile) int32 0 1 2
* time (time) datetime64[ns] 2010-03-16T12:00:00
XC (tile, j, i) float32 -111.6 -111.3 -110.9 ... 50.97 51.44 51.84
YC (tile, j, i) float32 -88.24 -88.38 -88.52 ... 67.62 67.53 67.47
... ...
Zp1 (k_p1) float32 0.0 -10.0 -20.0 ... -5.244e+03 -5.678e+03 -6.134e+03
Zu (k_u) float32 -10.0 -20.0 -30.0 ... -5.678e+03 -6.134e+03
Zl (k_l) float32 0.0 -10.0 -20.0 ... -4.834e+03 -5.244e+03 -5.678e+03
XC_bnds (tile, j, i, nb) float32 -115.0 -115.0 -107.9 ... 52.0 52.0 51.73
YC_bnds (tile, j, i, nb) float32 -88.18 -88.32 -88.3 ... 67.4 67.5 67.56
Z_bnds (k, nv) float32 0.0 -10.0 -10.0 ... -5.678e+03 -6.134e+03
Dimensions without coordinates: nb, nv
Data variables: (12/23)
SSH (time, tile, j, i) float32 nan nan nan nan nan ... nan nan nan nan
THETA (time, k, tile, j, i) float32 nan nan nan nan ... nan nan nan nan
CS (tile, j, i) float32 0.06158 0.06675 0.07293 ... 0.9052 0.9424
SN (tile, j, i) float32 -0.9981 -0.9978 -0.9973 ... -0.4251 -0.3344
rA (tile, j, i) float32 3.623e+08 3.633e+08 ... 3.71e+08 2.126e+08
dxG (tile, j_g, i) float32 1.558e+04 1.559e+04 ... 2.133e+04 1.787e+04
... ...
hFacC (k, tile, j, i) float32 0.0 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0 0.0
hFacW (k, tile, j, i_g) float32 0.0 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
hFacS (k, tile, j_g, i) float32 0.0 0.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
maskC (k, tile, j, i) bool False False False False ... False False False
maskW (k, tile, j, i_g) bool False False False ... False False False
maskS (k, tile, j_g, i) bool False False False ... False False False
Attributes:
long_name: Dynamic sea surface height anomaly
units: m
coverage_content_type: modelResult
standard_name: sea_surface_height_above_geoid
comment: Dynamic sea surface height anomaly above the geoi...
valid_min: -1.8805772066116333
valid_max: 1.4207719564437866- i: 90
- j: 90
- tile: 3
- time: 1
- k: 50
- i_g: 90
- j_g: 90
- k_u: 50
- k_l: 50
- k_p1: 51
- nb: 4
- nv: 2
- i(i)int320 1 2 3 4 5 6 ... 84 85 86 87 88 89
- axis :
- X
- long_name :
- grid index in x for variables at tracer and 'v' locations
- swap_dim :
- XC
- comment :
- In the Arakawa C-grid system, tracer (e.g., THETA) and 'v' variables (e.g., VVEL) have the same x coordinate on the model grid.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j(j)int320 1 2 3 4 5 6 ... 84 85 86 87 88 89
- axis :
- Y
- long_name :
- grid index in y for variables at tracer and 'u' locations
- swap_dim :
- YC
- comment :
- In the Arakawa C-grid system, tracer (e.g., THETA) and 'u' variables (e.g., UVEL) have the same y coordinate on the model grid.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - tile(tile)int320 1 2
- long_name :
- lat-lon-cap tile index
- comment :
- The ECCO V4 horizontal model grid is divided into 13 tiles of 90x90 cells for convenience.
- coverage_content_type :
- coordinate
array([0, 1, 2])
- time(time)datetime64[ns]2010-03-16T12:00:00
- long_name :
- center time of averaging period
- axis :
- T
- bounds :
- time_bnds
- coverage_content_type :
- coordinate
- standard_name :
- time
array(['2010-03-16T12:00:00.000000000'], dtype='datetime64[ns]')
- XC(tile, j, i)float32-111.6 -111.3 ... 51.44 51.84
- long_name :
- longitude of tracer grid cell center
- units :
- degrees_east
- coordinate :
- YC XC
- bounds :
- XC_bnds
- comment :
- nonuniform grid spacing
- coverage_content_type :
- coordinate
- standard_name :
- longitude
array([[[-111.60647 , -111.303 , -110.94285 , ..., 64.791115, 64.80521 , 64.81917 ], [-104.8196 , -103.928444, -102.87706 , ..., 64.36745 , 64.41012 , 64.4524 ], [ -98.198784, -96.788055, -95.14185 , ..., 63.936497, 64.008224, 64.0793 ], ..., [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ]], [[ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], ... [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ]], [[ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], ..., [ -37.730072, -37.17829 , -36.597565, ..., 50.597565, 51.17829 , 51.730072], [ -37.771988, -37.291943, -36.764027, ..., 50.764027, 51.291943, 51.771988], [ -37.837925, -37.44421 , -36.968143, ..., 50.968143, 51.44421 , 51.837925]]], dtype=float32) - YC(tile, j, i)float32-88.24 -88.38 ... 67.53 67.47
- long_name :
- latitude of tracer grid cell center
- units :
- degrees_north
- coordinate :
- YC XC
- bounds :
- YC_bnds
- comment :
- nonuniform grid spacing
- coverage_content_type :
- coordinate
- standard_name :
- latitude
array([[[-88.24259 , -88.382515 , -88.52242 , ..., -80.57383 , -80.50492 , -80.43992 ], [-88.21676 , -88.354485 , -88.49178 , ..., -80.56925 , -80.50038 , -80.43542 ], [-88.164764 , -88.29827 , -88.43065 , ..., -80.55984 , -80.49105 , -80.42617 ], ..., [-58.321964 , -58.321964 , -58.321964 , ..., -58.321964 , -58.321964 , -58.321964 ], [-57.79962 , -57.79962 , -57.79962 , ..., -57.79962 , -57.79962 , -57.79962 ], [-57.271408 , -57.271408 , -57.271408 , ..., -57.271408 , -57.271408 , -57.271408 ]], [[-56.73891 , -56.73891 , -56.73891 , ..., -56.73891 , -56.73891 , -56.73891 ], [-56.2021 , -56.2021 , -56.2021 , ..., -56.2021 , -56.2021 , -56.2021 ], [-55.65936 , -55.65936 , -55.65936 , ..., -55.65936 , -55.65936 , -55.65936 ], ... [ 7.5699615, 7.5699615, 7.5699615, ..., 7.5699615, 7.5699615, 7.5699615], [ 8.516253 , 8.516253 , 8.516253 , ..., 8.516253 , 8.516253 , 8.516253 ], [ 9.482398 , 9.482398 , 9.482398 , ..., 9.482398 , 9.482398 , 9.482398 ]], [[ 10.458642 , 10.458642 , 10.458642 , ..., 10.458642 , 10.458642 , 10.458642 ], [ 11.438585 , 11.438585 , 11.438585 , ..., 11.438585 , 11.438585 , 11.438585 ], [ 12.418467 , 12.418467 , 12.418467 , ..., 12.418467 , 12.418467 , 12.418467 ], ..., [ 67.16407 , 67.19428 , 67.24966 , ..., 67.24966 , 67.19428 , 67.16407 ], [ 67.33552 , 67.37607 , 67.445274 , ..., 67.445274 , 67.37607 , 67.33552 ], [ 67.47211 , 67.53387 , 67.62317 , ..., 67.62317 , 67.53387 , 67.47211 ]]], dtype=float32) - k(k)int320 1 2 3 4 5 6 ... 44 45 46 47 48 49
- axis :
- Z
- long_name :
- grid index in z for tracer variables
- swap_dim :
- Z
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - Z(k)float32-5.0 -15.0 ... -5.906e+03
- long_name :
- depth of tracer grid cell center
- units :
- m
- positive :
- up
- bounds :
- Z_bnds
- comment :
- Non-uniform vertical spacing.
- coverage_content_type :
- coordinate
- standard_name :
- depth
array([-5.000000e+00, -1.500000e+01, -2.500000e+01, -3.500000e+01, -4.500000e+01, -5.500000e+01, -6.500000e+01, -7.500500e+01, -8.502500e+01, -9.509500e+01, -1.053100e+02, -1.158700e+02, -1.271500e+02, -1.397400e+02, -1.544700e+02, -1.724000e+02, -1.947350e+02, -2.227100e+02, -2.574700e+02, -2.999300e+02, -3.506800e+02, -4.099300e+02, -4.774700e+02, -5.527100e+02, -6.347350e+02, -7.224000e+02, -8.144700e+02, -9.097400e+02, -1.007155e+03, -1.105905e+03, -1.205535e+03, -1.306205e+03, -1.409150e+03, -1.517095e+03, -1.634175e+03, -1.765135e+03, -1.914150e+03, -2.084035e+03, -2.276225e+03, -2.491250e+03, -2.729250e+03, -2.990250e+03, -3.274250e+03, -3.581250e+03, -3.911250e+03, -4.264250e+03, -4.640250e+03, -5.039250e+03, -5.461250e+03, -5.906250e+03], dtype=float32) - i_g(i_g)int320 1 2 3 4 5 6 ... 84 85 86 87 88 89
- axis :
- X
- long_name :
- grid index in x for variables at 'u' and 'g' locations
- c_grid_axis_shift :
- -0.5
- swap_dim :
- XG
- comment :
- In the Arakawa C-grid system, 'u' (e.g., UVEL) and 'g' variables (e.g., XG) have the same x coordinate on the model grid.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j_g(j_g)int320 1 2 3 4 5 6 ... 84 85 86 87 88 89
- axis :
- Y
- long_name :
- grid index in y for variables at 'v' and 'g' locations
- c_grid_axis_shift :
- -0.5
- swap_dim :
- YG
- comment :
- In the Arakawa C-grid system, 'v' (e.g., VVEL) and 'g' variables (e.g., XG) have the same y coordinate.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - k_u(k_u)int320 1 2 3 4 5 6 ... 44 45 46 47 48 49
- axis :
- Z
- long_name :
- grid index in z corresponding to the bottom face of tracer grid cells ('w' locations)
- c_grid_axis_shift :
- 0.5
- swap_dim :
- Zu
- comment :
- First index corresponds to the bottom face of the uppermost tracer grid cell. The use of 'u' in the variable name follows the MITgcm convention for naming the bottom face of ocean tracer grid cells.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_l(k_l)int320 1 2 3 4 5 6 ... 44 45 46 47 48 49
- axis :
- Z
- long_name :
- grid index in z corresponding to the top face of tracer grid cells ('w' locations)
- c_grid_axis_shift :
- -0.5
- swap_dim :
- Zl
- comment :
- First index corresponds to the top face of the uppermost tracer grid cell. The use of 'l' in the variable name follows the MITgcm convention for naming the top face of ocean tracer grid cells.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49]) - k_p1(k_p1)int320 1 2 3 4 5 6 ... 45 46 47 48 49 50
- axis :
- Z
- long_name :
- grid index in z for variables at 'w' locations
- c_grid_axis_shift :
- [-0.5 0.5]
- swap_dim :
- Zp1
- comment :
- Includes top of uppermost model tracer cell (k_p1=0) and bottom of lowermost tracer cell (k_p1=51).
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50]) - XG(tile, j_g, i_g)float32-115.0 -115.0 ... 51.12 51.59
- long_name :
- longitude of 'southwest' corner of tracer grid cell
- units :
- degrees_east
- coordinate :
- YG XG
- comment :
- Nonuniform grid spacing. Note: 'southwest' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm dcoumentation for details.
- coverage_content_type :
- coordinate
- standard_name :
- longitude
array([[[-115. , -115. , -115. , ..., 65. , 65. , 65. ], [-108.47213 , -107.915245, -107.26045 , ..., 64.56598 , 64.59443 , 64.62261 ], [-101.9954 , -100.91797 , -99.66073 , ..., 64.12406 , 64.18147 , 64.23835 ], ..., [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ]], [[ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], ... [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ]], [[ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], [ -38. , -37. , -36. , ..., 49. , 50. , 51. ], ..., [ -38. , -37.42438 , -36.831715, ..., 50.211548, 50.831715, 51.42438 ], [ -38. , -37.494156, -36.95921 , ..., 50.38373 , 50.95921 , 51.494156], [ -38. , -37.588932, -37.118958, ..., 50.58833 , 51.118958, 51.588932]]], dtype=float32) - YG(tile, j_g, i_g)float32-88.18 -88.32 ... 67.49 67.43
- long_name :
- latitude of 'southwest' corner of tracer grid cell
- units :
- degrees_north
- comment :
- Nonuniform grid spacing. Note: 'southwest' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm dcoumentation for details.
- coverage_content_type :
- coordinate
- standard_name :
- latitude
array([[[-88.17569 , -88.31587 , -88.45609 , ..., -80.610275, -80.53945 , -80.47247 ], [-88.16335 , -88.30251 , -88.44153 , ..., -80.608 , -80.5372 , -80.470245], [-88.12586 , -88.26202 , -88.397514, ..., -80.601006, -80.530266, -80.46337 ], ..., [-58.58002 , -58.58002 , -58.58002 , ..., -58.58002 , -58.58002 , -58.58002 ], [-58.06183 , -58.06183 , -58.06183 , ..., -58.06183 , -58.06183 , -58.06183 ], [-57.53605 , -57.53605 , -57.53605 , ..., -57.53605 , -57.53605 , -57.53605 ]], [[-57.005695, -57.005695, -57.005695, ..., -57.005695, -57.005695, -57.005695], [-56.471046, -56.471046, -56.471046, ..., -56.471046, -56.471046, -56.471046], [-55.931786, -55.931786, -55.931786, ..., -55.931786, -55.931786, -55.931786], ... [ 7.108203, 7.108203, 7.108203, ..., 7.108203, 7.108203, 7.108203], [ 8.039881, 8.039881, 8.039881, ..., 8.039881, 8.039881, 8.039881], [ 8.997536, 8.997536, 8.997536, ..., 8.997536, 8.997536, 8.997536]], [[ 9.96973 , 9.96973 , 9.96973 , ..., 9.96973 , 9.96973 , 9.96973 ], [ 10.948437, 10.948437, 10.948437, ..., 10.948437, 10.948437, 10.948437], [ 11.928692, 11.928692, 11.928692, ..., 11.928692, 11.928692, 11.928692], ..., [ 67.06448 , 67.07818 , 67.11726 , ..., 67.17736 , 67.11726 , 67.07818 ], [ 67.24895 , 67.26691 , 67.31608 , ..., 67.38828 , 67.31608 , 67.26691 ], [ 67.40169 , 67.42869 , 67.49435 , ..., 67.58286 , 67.49435 , 67.42869 ]]], dtype=float32) - Zp1(k_p1)float320.0 -10.0 ... -5.678e+03 -6.134e+03
- long_name :
- depth of top/bottom face of tracer grid cell
- units :
- m
- positive :
- up
- comment :
- Contains one element more than the number of vertical layers. First element is 0m, the depth of the top face of the uppermost grid cell. Last element is the depth of the bottom face of the deepest grid cell.
- coverage_content_type :
- coordinate
- standard_name :
- depth
array([ 0. , -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. , -6134.5 ], dtype=float32) - Zu(k_u)float32-10.0 -20.0 ... -6.134e+03
- long_name :
- depth of bottom face of tracer grid cell
- units :
- m
- positive :
- up
- comment :
- First element is -10m, the depth of the bottom face of the uppermost tracer grid cell. Last element is the depth of the bottom face of the deepest grid cell. The use of 'u' in the variable name follows the MITgcm convention for naming the bottom face of ocean tracer grid cells.
- coverage_content_type :
- coordinate
- standard_name :
- depth
array([ -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. , -6134.5 ], dtype=float32) - Zl(k_l)float320.0 -10.0 ... -5.244e+03 -5.678e+03
- long_name :
- depth of top face of tracer grid cell
- units :
- m
- positive :
- up
- comment :
- First element is 0m, the depth of the top face of the uppermost tracer grid cell (i.e., the ocean surface). Last element is the depth of the top face of the deepest grid cell. The use of 'l' in the variable name follows the MITgcm convention for naming the top face of ocean tracer grid cells.
- coverage_content_type :
- coordinate
- standard_name :
- depth
array([ 0. , -10. , -20. , -30. , -40. , -50. , -60. , -70. , -80.01, -90.04, -100.15, -110.47, -121.27, -133.03, -146.45, -162.49, -182.31, -207.16, -238.26, -276.68, -323.18, -378.18, -441.68, -513.26, -592.16, -677.31, -767.49, -861.45, -958.03, -1056.28, -1155.53, -1255.54, -1356.87, -1461.43, -1572.76, -1695.59, -1834.68, -1993.62, -2174.45, -2378. , -2604.5 , -2854. , -3126.5 , -3422. , -3740.5 , -4082. , -4446.5 , -4834. , -5244.5 , -5678. ], dtype=float32) - XC_bnds(tile, j, i, nb)float32-115.0 -115.0 -107.9 ... 52.0 51.73
- comment :
- Bounds array follows CF conventions. XC_bnds[i,j,0] = 'southwest' corner (j-1, i-1), XC_bnds[i,j,1] = 'southeast' corner (j-1, i+1), XC_bnds[i,j,2] = 'northeast' corner (j+1, i+1), XC_bnds[i,j,3] = 'northwest' corner (j+1, i-1). Note: 'southwest', 'southeast', northwest', and 'northeast' do not correspond to geographic orientation but are used for convenience to describe the computational grid. See MITgcm dcoumentation for details.
- coverage_content_type :
- coordinate
- long_name :
- longitudes of tracer grid cell corners
array([[[[-115. , -115. , -107.915245, -108.47213 ], [-115. , -115. , -107.26045 , -107.915245], [-115. , -115. , -106.47914 , -107.26045 ], ..., [ 65. , 65. , 64.59443 , 64.56598 ], [ 65. , 65. , 64.62261 , 64.59443 ], [ 65. , 65. , 64.65055 , 64.62261 ]], [[-108.47213 , -107.915245, -100.91797 , -101.9954 ], [-107.915245, -107.26045 , -99.66073 , -100.91797 ], [-107.26045 , -106.47914 , -98.17517 , -99.66073 ], ..., [ 64.56598 , 64.59443 , 64.18147 , 64.12406 ], [ 64.59443 , 64.62261 , 64.23835 , 64.18147 ], [ 64.62261 , 64.65055 , 64.29474 , 64.23835 ]], [[-101.9954 , -100.91797 , -94.218956, -95.74128 ], [-100.91797 , -99.66073 , -92.46296 , -94.218956], [ -99.66073 , -98.17517 , -90.417885, -92.46296 ], ..., ... [ 50.211548, 50.831715, 50.95921 , 50.38373 ], [ 50.831715, 51.42438 , 51.494156, 50.95921 ], [ 51.42438 , 52. , 52. , 51.494156]], [[ -38. , -37.494156, -37.588932, -38. ], [ -37.494156, -36.95921 , -37.118958, -37.588932], [ -36.95921 , -36.38373 , -36.58833 , -37.118958], ..., [ 50.38373 , 50.95921 , 51.118958, 50.58833 ], [ 50.95921 , 51.494156, 51.588932, 51.118958], [ 51.494156, 52. , 52. , 51.588932]], [[ -38. , -37.588932, -37.734455, -38. ], [ -37.588932, -37.118958, -37.324615, -37.734455], [ -37.118958, -36.58833 , -36.832813, -37.324615], ..., [ 50.58833 , 51.118958, 51.324615, 50.832813], [ 51.118958, 51.588932, 51.734455, 51.324615], [ 51.588932, 52. , 52. , 51.734455]]]], dtype=float32) - YC_bnds(tile, j, i, nb)float32-88.18 -88.32 -88.3 ... 67.5 67.56
- comment :
- Bounds array follows CF conventions. YC_bnds[i,j,0] = 'southwest' corner (j-1, i-1), YC_bnds[i,j,1] = 'southeast' corner (j-1, i+1), YC_bnds[i,j,2] = 'northeast' corner (j+1, i+1), YC_bnds[i,j,3] = 'northwest' corner (j+1, i-1). Note: 'southwest', 'southeast', northwest', and 'northeast' do not correspond to geographic orientation but are used for convenience to describe the computational grid. See MITgcm dcoumentation for details.
- coverage_content_type :
- coordinate
- long_name :
- latitudes of tracer grid cell corners
array([[[[-88.17569 , -88.31587 , -88.30251 , -88.16335 ], [-88.31587 , -88.45609 , -88.44153 , -88.30251 ], [-88.45609 , -88.596344, -88.580345, -88.44153 ], ..., [-80.610275, -80.53945 , -80.5372 , -80.608 ], [-80.53945 , -80.47247 , -80.470245, -80.5372 ], [-80.47247 , -80.40948 , -80.40726 , -80.470245]], [[-88.16335 , -88.30251 , -88.26202 , -88.12586 ], [-88.30251 , -88.44153 , -88.397514, -88.26202 ], [-88.44153 , -88.580345, -88.53216 , -88.397514], ..., [-80.608 , -80.5372 , -80.530266, -80.601006], [-80.5372 , -80.470245, -80.46337 , -80.530266], [-80.470245, -80.40726 , -80.400444, -80.46337 ]], [[-88.12586 , -88.26202 , -88.194695, -88.06328 ], [-88.26202 , -88.397514, -88.32474 , -88.194695], [-88.397514, -88.53216 , -88.453026, -88.32474 ], ..., ... [ 67.17736 , 67.11726 , 67.31608 , 67.38828 ], [ 67.11726 , 67.07818 , 67.26691 , 67.31608 ], [ 67.07818 , 67.06448 , 67.24895 , 67.26691 ]], [[ 67.24895 , 67.26691 , 67.42869 , 67.40169 ], [ 67.26691 , 67.31608 , 67.49435 , 67.42869 ], [ 67.31608 , 67.38828 , 67.58286 , 67.49435 ], ..., [ 67.38828 , 67.31608 , 67.49435 , 67.58286 ], [ 67.31608 , 67.26691 , 67.42869 , 67.49435 ], [ 67.26691 , 67.24895 , 67.40169 , 67.42869 ]], [[ 67.40169 , 67.42869 , 67.560646, 67.5 ], [ 67.42869 , 67.49435 , 67.6532 , 67.560646], [ 67.49435 , 67.58286 , 67.76259 , 67.6532 ], ..., [ 67.58286 , 67.49435 , 67.6532 , 67.76259 ], [ 67.49435 , 67.42869 , 67.560646, 67.6532 ], [ 67.42869 , 67.40169 , 67.5 , 67.560646]]]], dtype=float32) - Z_bnds(k, nv)float320.0 -10.0 ... -5.678e+03 -6.134e+03
- comment :
- One pair of depths for each vertical level.
- coverage_content_type :
- coordinate
- long_name :
- depths of top and bottom faces of tracer grid cell
array([[ 0. , -10. ], [ -10. , -20. ], [ -20. , -30. ], [ -30. , -40. ], [ -40. , -50. ], [ -50. , -60. ], [ -60. , -70. ], [ -70. , -80.01 ], [ -80.01 , -90.04 ], [ -90.04 , -100.15 ], [ -100.15 , -110.47 ], [ -110.47 , -121.270004], [ -121.270004, -133.03 ], [ -133.03 , -146.45 ], [ -146.45 , -162.48999 ], [ -162.48999 , -182.31 ], [ -182.31 , -207.16 ], [ -207.16 , -238.26001 ], [ -238.26001 , -276.68 ], [ -276.68 , -323.18 ], ... [-1155.53 , -1255.54 ], [-1255.54 , -1356.87 ], [-1356.87 , -1461.4299 ], [-1461.4299 , -1572.7599 ], [-1572.7599 , -1695.5898 ], [-1695.5898 , -1834.6798 ], [-1834.6798 , -1993.6199 ], [-1993.6199 , -2174.45 ], [-2174.45 , -2378. ], [-2378. , -2604.5 ], [-2604.5 , -2854. ], [-2854. , -3126.5 ], [-3126.5 , -3422. ], [-3422. , -3740.5 ], [-3740.5 , -4082. ], [-4082. , -4446.5 ], [-4446.5 , -4834. ], [-4834. , -5244.5 ], [-5244.5 , -5678. ], [-5678. , -6134.5 ]], dtype=float32)
- SSH(time, tile, j, i)float32nan nan nan nan ... nan nan nan nan
- long_name :
- Dynamic sea surface height anomaly
- units :
- m
- coverage_content_type :
- modelResult
- standard_name :
- sea_surface_height_above_geoid
- comment :
- Dynamic sea surface height anomaly above the geoid, suitable for comparisons with altimetry sea surface height data products that apply the inverse barometer (IB) correction. Note: SSH is calculated by correcting model sea level anomaly ETAN for three effects: a) global mean steric sea level changes related to density changes in the Boussinesq volume-conserving model (Greatbatch correction, see sterGloH), b) the inverted barometer (IB) effect (see SSHIBC) and c) sea level displacement due to sea-ice and snow pressure loading (see sIceLoad). SSH can be compared with the similarly-named SSH variable in previous ECCO products that did not include atmospheric pressure loading (e.g., Version 4 Release 3). Use SSHNOIBC for comparisons with altimetry data products that do NOT apply the IB correction.
- valid_min :
- -1.8805772066116333
- valid_max :
- 1.4207719564437866
array([[[[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [-1.437773 , -1.4438914 , -1.4436442 , ..., -1.3392228 , -1.3288928 , -1.318716 ], [-1.409977 , -1.4160229 , -1.4151381 , ..., -1.3262419 , -1.3157413 , -1.3043153 ], [-1.3782076 , -1.3828527 , -1.3819864 , ..., -1.3157756 , -1.3051451 , -1.2920415 ]], [[-1.3424112 , -1.3456359 , -1.3465977 , ..., -1.304871 , -1.2944611 , -1.2798376 ], [-1.3084964 , -1.3112303 , -1.315978 , ..., -1.2911572 , -1.281555 , -1.2661443 ], [-1.2859204 , -1.2886494 , -1.2975372 , ..., -1.2729706 , -1.2647922 , -1.2497054 ], ... [ 0.18926093, 0.17679782, 0.16287906, ..., nan, 0.531225 , 0.55789405], [ 0.17079698, 0.15768045, 0.14356998, ..., nan, 0.49427482, 0.5214947 ], [ 0.15235202, 0.1392552 , 0.12553291, ..., nan, nan, 0.48885915]], [[ 0.13727026, 0.12395398, 0.11061972, ..., nan, nan, 0.46835414], [ 0.12681508, 0.11316036, 0.09986742, ..., 0.38901928, nan, 0.45423734], [ 0.1223011 , 0.10851809, 0.0957661 , ..., 0.4008969 , 0.41220444, 0.44959423], ..., [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan]]]], dtype=float32) - THETA(time, k, tile, j, i)float32nan nan nan nan ... nan nan nan nan
- long_name :
- Potential temperature
- units :
- degree_C
- coverage_content_type :
- modelResult
- standard_name :
- sea_water_potential_temperature
- comment :
- Sea water potential temperature is the temperature a parcel of sea water would have if moved adiabatically to sea level pressure. Note: the equation of state is a modified UNESCO formula by Jackett and McDougall (1995), which uses the model variable potential temperature as input assuming a horizontally and temporally constant pressure of $p_0=-g \rho_{0} z$.
- valid_min :
- -2.2909388542175293
- valid_max :
- 36.032955169677734
array([[[[[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [ 2.4149065 , 2.4036155 , 2.3684697 , ..., 1.6430806 , 1.6572165 , 1.669287 ], [ 2.5654478 , 2.5598085 , 2.526788 , ..., 1.7637222 , 1.7874517 , 1.8169286 ], [ 2.697345 , 2.69822 , 2.6639824 , ..., 1.8744359 , 1.9126662 , 1.9653959 ]], [[ 2.797386 , 2.7955499 , 2.748563 , ..., 1.9828016 , 2.0356798 , 2.1142213 ], [ 2.8660307 , 2.854989 , 2.7869945 , ..., 2.099932 , 2.1638968 , 2.2642674 ], [ 2.9065216 , 2.8897736 , 2.7960505 , ..., 2.2311447 , 2.2983036 , 2.408241 ], ... [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan]], [[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan]]]]], dtype=float32) - CS(tile, j, i)float320.06158 0.06675 ... 0.9052 0.9424
- long_name :
- cosine of tracer grid cell orientation vs geographical north
- units :
- 1
- coordinate :
- YC XC
- coverage_content_type :
- modelResult
- comment :
- CS and SN are required to calculate the geographic (meridional, zonal) components of vectors on the curvilinear model grid. Note: for vector R with components R_x and R_y: R_{east} = CS R_x - SN R_y. R_{north} = SN R_x + CS R_y
array([[[0.06157813, 0.06675376, 0.07293081, ..., 0.03294202, 0.03476389, 0.03689073], [0.18319611, 0.19806239, 0.21563956, ..., 0.09903675, 0.10446203, 0.1107849 ], [0.29926026, 0.32193854, 0.34829748, ..., 0.16525467, 0.17413175, 0.18444225], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ..., [0.9970846 , 0.9799519 , 0.96124434, ..., 0.96124434, 0.9799519 , 0.9970846 ], [0.99148935, 0.95813316, 0.9350622 , ..., 0.9350622 , 0.95813316, 0.99148935], [0.9424238 , 0.9051672 , 0.8916124 , ..., 0.8916124 , 0.9051672 , 0.9424238 ]]], dtype=float32) - SN(tile, j, i)float32-0.9981 -0.9978 ... -0.4251 -0.3344
- long_name :
- sine of tracer grid cell orientation vs geographical north
- units :
- 1
- coordinate :
- YC XC
- coverage_content_type :
- modelResult
- comment :
- CS and SN are required to calculate the geographic (meridional, zonal) components of vectors on the curvilinear model grid. Note: for vector R with components R_x and R_y in local grid directions x and y, the geographical eastward component R_{east} = CS R_x - SN R_y. The geographical northward component R_{north} = SN R_x + CS R_y.
array([[[-9.9810225e-01, -9.9776947e-01, -9.9733698e-01, ..., 9.9945724e-01, 9.9939555e-01, 9.9931931e-01], [-9.8307639e-01, -9.8018944e-01, -9.7647303e-01, ..., 9.9508375e-01, 9.9452889e-01, 9.9384439e-01], [-9.5417154e-01, -9.4676059e-01, -9.3738407e-01, ..., 9.8625094e-01, 9.8472238e-01, 9.8284334e-01], ..., [ 1.3431713e-14, -6.6167042e-15, -6.8150090e-15, ..., 6.8150090e-15, 6.6167042e-15, -1.3431713e-14], [ 1.3431713e-14, -1.3431713e-14, -0.0000000e+00, ..., -0.0000000e+00, 1.3431713e-14, -1.3431713e-14], [-0.0000000e+00, 6.5240599e-15, -6.5240599e-15, ..., 6.5240599e-15, -6.5240599e-15, -0.0000000e+00]], [[-0.0000000e+00, 6.5240599e-15, -6.5240599e-15, ..., 6.5240599e-15, -6.5240599e-15, -0.0000000e+00], [-0.0000000e+00, -0.0000000e+00, 6.3420986e-15, ..., -6.3420986e-15, -0.0000000e+00, -0.0000000e+00], [-1.2508176e-14, -0.0000000e+00, 6.3420986e-15, ..., -6.3420986e-15, -0.0000000e+00, 1.2508176e-14], ... [ 4.4752880e-16, -1.3445239e-15, 8.9699497e-16, ..., -8.9699497e-16, 1.3445239e-15, -4.4752880e-16], [ 8.9924353e-16, -1.7962386e-15, 1.7962386e-15, ..., -1.7962386e-15, 1.7962386e-15, -8.9924353e-16], [-2.5525710e-18, -8.9924353e-16, 1.8010396e-15, ..., -1.8010396e-15, 8.9924353e-16, 2.5525710e-18]], [[-1.8064405e-15, 9.0464432e-16, -2.8481762e-18, ..., 2.8481762e-18, -9.0464432e-16, 1.8064405e-15], [-9.0464432e-16, -3.1371137e-18, 3.1371137e-18, ..., -3.1371137e-18, 3.1371137e-18, 9.0464432e-16], [-9.1120342e-16, 9.1462544e-16, -3.4220292e-18, ..., 3.4220292e-18, -9.1462544e-16, 9.1120342e-16], ..., [ 7.6303728e-02, 1.9923417e-01, 2.7569798e-01, ..., -2.7569798e-01, -1.9923417e-01, -7.6303728e-02], [ 1.3018784e-01, 2.8632292e-01, 3.5448366e-01, ..., -3.5448366e-01, -2.8632292e-01, -1.3018784e-01], [ 3.3442083e-01, 4.2505559e-01, 4.5279941e-01, ..., -4.5279941e-01, -4.2505559e-01, -3.3442083e-01]]], dtype=float32) - rA(tile, j, i)float323.623e+08 3.633e+08 ... 2.126e+08
- long_name :
- area of tracer grid cell
- units :
- m2
- coordinate :
- YC XC
- coverage_content_type :
- modelResult
- standard_name :
- cell_area
array([[[3.6225645e+08, 3.6330096e+08, 3.6425389e+08, ..., 6.0276308e+07, 5.3553676e+07, 4.7093872e+07], [3.6961037e+08, 3.7067478e+08, 3.7164586e+08, ..., 6.1906760e+07, 5.5057284e+07, 4.8475600e+07], [3.7716797e+08, 3.7825142e+08, 3.7923990e+08, ..., 6.4021868e+07, 5.7052416e+07, 5.0355492e+07], ..., [3.3636692e+09, 3.3636692e+09, 3.3636692e+09, ..., 3.3636692e+09, 3.3636692e+09, 3.3636692e+09], [3.4631982e+09, 3.4631982e+09, 3.4631982e+09, ..., 3.4631982e+09, 3.4631982e+09, 3.4631982e+09], [3.5442668e+09, 3.5442668e+09, 3.5442668e+09, ..., 3.5442668e+09, 3.5442668e+09, 3.5442668e+09]], [[3.6245120e+09, 3.6245120e+09, 3.6245120e+09, ..., 3.6245120e+09, 3.6245120e+09, 3.6245120e+09], [3.7078205e+09, 3.7078205e+09, 3.7078205e+09, ..., 3.7078205e+09, 3.7078205e+09, 3.7078205e+09], [3.8142915e+09, 3.8142915e+09, 3.8142915e+09, ..., 3.8142915e+09, 3.8142915e+09, 3.8142915e+09], ... [1.1415347e+10, 1.1415347e+10, 1.1415347e+10, ..., 1.1415347e+10, 1.1415347e+10, 1.1415347e+10], [1.1706299e+10, 1.1706299e+10, 1.1706299e+10, ..., 1.1706299e+10, 1.1706299e+10, 1.1706299e+10], [1.1852359e+10, 1.1852359e+10, 1.1852359e+10, ..., 1.1852359e+10, 1.1852359e+10, 1.1852359e+10]], [[1.1896091e+10, 1.1896091e+10, 1.1896091e+10, ..., 1.1896091e+10, 1.1896091e+10, 1.1896091e+10], [1.1875574e+10, 1.1875574e+10, 1.1875574e+10, ..., 1.1875574e+10, 1.1875574e+10, 1.1875574e+10], [1.1817668e+10, 1.1817668e+10, 1.1817668e+10, ..., 1.1817668e+10, 1.1817668e+10, 1.1817668e+10], ..., [4.8800346e+08, 5.4557037e+08, 6.3389197e+08, ..., 6.3389197e+08, 5.4557037e+08, 4.8800346e+08], [3.5101645e+08, 4.4306134e+08, 5.5967296e+08, ..., 5.5967296e+08, 4.4306134e+08, 3.5101645e+08], [2.1263387e+08, 3.7097568e+08, 5.1378093e+08, ..., 5.1378093e+08, 3.7097568e+08, 2.1263387e+08]]], dtype=float32) - dxG(tile, j_g, i)float321.558e+04 1.559e+04 ... 1.787e+04
- long_name :
- distance between 'southwest' and 'southeast' corners of the tracer grid cell
- units :
- m
- coordinate :
- YG XC
- coverage_content_type :
- modelResult
- comment :
- Alternatively, the length of 'south' side of tracer grid cell. Note: 'south', 'southwest', and 'southeast' do not correspond to geographic orientation but are used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[ 15584.907 , 15589.316 , 15593.201 , ..., 7873.7295, 7446.052 , 7004.153 ], [ 15588.651 , 15593.048 , 15596.92 , ..., 7888.4585, 7461.8286, 7021.1333], [ 15600.185 , 15604.543 , 15608.381 , ..., 7933.642 , 7510.2056, 7073.169 ], ..., [ 57957.625 , 57957.625 , 57957.625 , ..., 57957.625 , 57957.625 , 57957.625 ], [ 58813.305 , 58813.305 , 58813.305 , ..., 58813.305 , 58813.305 , 58813.305 ], [ 59676.61 , 59676.61 , 59676.61 , ..., 59676.61 , 59676.61 , 59676.61 ]], [[ 60542.324 , 60542.324 , 60542.324 , ..., 60542.324 , 60542.324 , 60542.324 ], [ 61409.805 , 61409.805 , 61409.805 , ..., 61409.805 , 61409.805 , 61409.805 ], [ 62279.344 , 62279.344 , 62279.344 , ..., 62279.344 , 62279.344 , 62279.344 ], ... [110322.99 , 110322.99 , 110322.99 , ..., 110322.99 , 110322.99 , 110322.99 ], [110084.7 , 110084.7 , 110084.7 , ..., 110084.7 , 110084.7 , 110084.7 ], [109809.445 , 109809.445 , 109809.445 , ..., 109809.445 , 109809.445 , 109809.445 ]], [[109498.625 , 109498.625 , 109498.625 , ..., 109498.625 , 109498.625 , 109498.625 ], [109153.875 , 109153.875 , 109153.875 , ..., 109153.875 , 109153.875 , 109153.875 ], [108776.664 , 108776.664 , 108776.664 , ..., 108776.664 , 108776.664 , 108776.664 ], ..., [ 24995.342 , 26019.037 , 27603.951 , ..., 27603.951 , 26019.037 , 24995.342 ], [ 21862.79 , 23614.71 , 25916.31 , ..., 25916.31 , 23614.71 , 21862.79 ], [ 17873.049 , 21326.564 , 24596.092 , ..., 24596.092 , 21326.564 , 17873.049 ]]], dtype=float32) - dyG(tile, j, i_g)float322.321e+04 2.327e+04 ... 1.597e+04
- long_name :
- distance between 'southwest' and 'northwest' corners of the tracer grid cell
- units :
- m
- coordinate :
- YC XG
- coverage_content_type :
- modelResult
- comment :
- Alternatively, the length of 'west' side of tracer grid cell. Note: 'west, 'southwest', and 'northwest' do not correspond to geographic orientation but are used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[ 23210.262 , 23273.26 , 23331.125 , ..., 7878.489 , 7417.633 , 6951.1924], [ 23669.895 , 23734.096 , 23793.066 , ..., 8059.287 , 7591.293 , 7117.8667], [ 24129.756 , 24195.113 , 24255.148 , ..., 8267.303 , 7794.305 , 7316.3003], ..., [ 57611.023 , 57611.023 , 57611.023 , ..., 57611.023 , 57611.023 , 57611.023 ], [ 58455.164 , 58455.164 , 58455.164 , ..., 58455.164 , 58455.164 , 58455.164 ], [ 58963.117 , 58963.117 , 58963.117 , ..., 58963.117 , 58963.117 , 58963.117 ]], [[ 59441.125 , 59441.125 , 59441.125 , ..., 59441.125 , 59441.125 , 59441.125 ], [ 59953.41 , 59953.41 , 59953.41 , ..., 59953.41 , 59953.41 , 59953.41 ], [ 60816.504 , 60816.504 , 60816.504 , ..., 60816.504 , 60816.504 , 60816.504 ], ... [103581.64 , 103581.64 , 103581.64 , ..., 103581.64 , 103581.64 , 103581.64 ], [106469.66 , 106469.66 , 106469.66 , ..., 106469.66 , 106469.66 , 106469.66 ], [108086.08 , 108086.08 , 108086.08 , ..., 108086.08 , 108086.08 , 108086.08 ]], [[108810.11 , 108810.11 , 108810.11 , ..., 108810.11 , 108810.11 , 108810.11 ], [108982.29 , 108982.29 , 108982.29 , ..., 108982.29 , 108982.29 , 108982.29 ], [108843.305 , 108843.305 , 108843.305 , ..., 108843.305 , 108843.305 , 108843.305 ], ..., [ 20508.13 , 21199.861 , 22779.795 , ..., 24591.26 , 22779.795 , 21199.861 ], [ 16981.354 , 18448.256 , 20970.678 , ..., 23327.838 , 20970.678 , 18448.256 ], [ 10930.409 , 15965.378 , 19713.574 , ..., 22499.89 , 19713.574 , 15965.378 ]]], dtype=float32) - Depth(tile, j, i)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- long_name :
- model seafloor depth below ocean surface at rest
- units :
- m
- coordinate :
- XC YC
- coverage_content_type :
- modelResult
- standard_name :
- sea_floor_depth_below_geoid
- comment :
- Model sea surface height (SSH) of 0m corresponds to an ocean surface at rest relative to the geoid. Depth corresponds to seafloor depth below geoid. Note: the MITgcm used by ECCO V4r4 implements 'partial cells' so the actual model seafloor depth may differ from the seafloor depth provided by the input bathymetry file.
array([[[ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [2977.8345 , 2503.9146 , 2562.9644 , ..., 5431.1567 , 5356.7544 , 5331.2 ], [3125.5176 , 2908.5 , 3046.7676 , ..., 5440.6733 , 5383.5386 , 5332.443 ], [3203.0798 , 3283.116 , 3312.7979 , ..., 5455.109 , 5433.769 , 5367.7656 ]], [[3284.1084 , 3485.7 , 3485.7 , ..., 5455.109 , 5444.462 , 5387.942 ], [3350.4321 , 3403.6208 , 3485.7 , ..., 5367.847 , 5379.0474 , 5359.3574 ], [3324.7493 , 3126.5 , 2243.637 , ..., 5244.5 , 5244.5 , 5196.211 ], ... [3865.2014 , 3902.5327 , 3985.4739 , ..., 0. , 2836.1733 , 4708.254 ], [3931.512 , 4203.5933 , 4347.109 , ..., 0. , 645.98663, 4068.4116 ], [4220.781 , 4446.5 , 4678.801 , ..., 0. , 0. , 1310.237 ]], [[4658.681 , 4820.5703 , 5014.177 , ..., 0. , 0. , 207.16 ], [4950.0605 , 5099.767 , 5237.27 , ..., 1993.62 , 0. , 219.7829 ], [5244.5 , 5493.2476 , 5633.637 , ..., 2164.8904 , 2164.8904 , 880.766 ], ..., [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ], [ 0. , 0. , 0. , ..., 0. , 0. , 0. ]]], dtype=float32) - rAz(tile, j_g, i_g)float321.799e+08 1.805e+08 ... 3.353e+08
- long_name :
- area of vorticity 'g' grid cell
- units :
- m2
- coordinate :
- YG XG
- coverage_content_type :
- modelResult
- standard_name :
- cell_area
- comment :
- Vorticity cells are staggered in space relative to tracer cells, nominally situated on tracer cell corners. Vorticity cell (i,j) is located at the 'southwest' corner of tracer grid cell (i, j). Note: 'southwest' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[1.7994426e+08, 1.8048699e+08, 1.8098384e+08, ..., 3.1679438e+07, 2.8274864e+07, 2.4993222e+07], [3.6534634e+08, 3.6644758e+08, 3.6745574e+08, ..., 6.4516980e+07, 5.7609388e+07, 5.0951208e+07], [3.7278982e+08, 3.7391142e+08, 3.7493824e+08, ..., 6.6447016e+07, 5.9413688e+07, 5.2634360e+07], ..., [3.3124155e+09, 3.3124155e+09, 3.3124155e+09, ..., 3.3124155e+09, 3.3124155e+09, 3.3124155e+09], [3.4155474e+09, 3.4155474e+09, 3.4155474e+09, ..., 3.4155474e+09, 3.4155474e+09, 3.4155474e+09], [3.5045770e+09, 3.5045770e+09, 3.5045770e+09, ..., 3.5045770e+09, 3.5045770e+09, 3.5045770e+09]], [[3.5842450e+09, 3.5842450e+09, 3.5842450e+09, ..., 3.5842450e+09, 3.5842450e+09, 3.5842450e+09], [3.6650668e+09, 3.6650668e+09, 3.6650668e+09, ..., 3.6650668e+09, 3.6650668e+09, 3.6650668e+09], [3.7580419e+09, 3.7580419e+09, 3.7580419e+09, ..., 3.7580419e+09, 3.7580419e+09, 3.7580419e+09], ... [1.1204871e+10, 1.1204871e+10, 1.1204871e+10, ..., 1.1204871e+10, 1.1204871e+10, 1.1204871e+10], [1.1581381e+10, 1.1581381e+10, 1.1581381e+10, ..., 1.1581381e+10, 1.1581381e+10, 1.1581381e+10], [1.1794822e+10, 1.1794822e+10, 1.1794822e+10, ..., 1.1794822e+10, 1.1794822e+10, 1.1794822e+10]], [[1.1884406e+10, 1.1884406e+10, 1.1884406e+10, ..., 1.1884406e+10, 1.1884406e+10, 1.1884406e+10], [1.1891903e+10, 1.1891903e+10, 1.1891903e+10, ..., 1.1891903e+10, 1.1891903e+10, 1.1891903e+10], [1.1850071e+10, 1.1850071e+10, 1.1850071e+10, ..., 1.1850071e+10, 1.1850071e+10, 1.1850071e+10], ..., [5.4691232e+08, 5.7209101e+08, 6.3671290e+08, ..., 7.2144787e+08, 6.3671290e+08, 5.7209101e+08], [4.0979427e+08, 4.5013763e+08, 5.4163482e+08, ..., 6.4757312e+08, 5.4163482e+08, 4.5013763e+08], [2.5678022e+08, 3.3529254e+08, 4.6752506e+08, ..., 5.9457242e+08, 4.6752506e+08, 3.3529254e+08]]], dtype=float32) - dxC(tile, j, i_g)float321.558e+04 1.559e+04 ... 1.812e+04
- long_name :
- distance between centers of adjacent tracer grid cells in the 'x' direction
- units :
- m
- coordinate :
- YC XG
- coverage_content_type :
- modelResult
- comment :
- Alternatively, the length of 'north' side of vorticity grid cells. Note: 'north' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[ 15583.418 , 15588.104 , 15592.241 , ..., 8085.6636, 7665.4624, 7230.907 ], [ 15591.017 , 15595.676 , 15599.79 , ..., 8114.4873, 7696.267 , 7263.974 ], [ 15606.636 , 15611.241 , 15615.306 , ..., 8173.435 , 7759.222 , 7331.496 ], ..., [ 58384.344 , 58384.344 , 58384.344 , ..., 58384.344 , 58384.344 , 58384.344 ], [ 59244.46 , 59244.46 , 59244.46 , ..., 59244.46 , 59244.46 , 59244.46 ], [ 60109.234 , 60109.234 , 60109.234 , ..., 60109.234 , 60109.234 , 60109.234 ]], [[ 60975.85 , 60975.85 , 60975.85 , ..., 60975.85 , 60975.85 , 60975.85 ], [ 61844.16 , 61844.16 , 61844.16 , ..., 61844.16 , 61844.16 , 61844.16 ], [ 62716.535 , 62716.535 , 62716.535 , ..., 62716.535 , 62716.535 , 62716.535 ], ... [110208.53 , 110208.53 , 110208.53 , ..., 110208.53 , 110208.53 , 110208.53 ], [109951.62 , 109951.62 , 109951.62 , ..., 109951.62 , 109951.62 , 109951.62 ], [109658.375 , 109658.375 , 109658.375 , ..., 109658.375 , 109658.375 , 109658.375 ]], [[109330.39 , 109330.39 , 109330.39 , ..., 109330.39 , 109330.39 , 109330.39 ], [108969.25 , 108969.25 , 108969.25 , ..., 108969.25 , 108969.25 , 108969.25 ], [108576.26 , 108576.26 , 108576.26 , ..., 108576.26 , 108576.26 , 108576.26 ], ..., [ 23317.729 , 24046.441 , 25751.87 , ..., 27725.76 , 25751.87 , 24046.441 ], [ 19590.729 , 21062.102 , 23828.547 , ..., 26457.6 , 23828.547 , 21062.102 ], [ 14067.07 , 18116.71 , 22498.332 , ..., 25686.383 , 22498.332 , 18116.71 ]]], dtype=float32) - dyC(tile, j_g, i)float321.156e+04 1.159e+04 ... 1.546e+04
- long_name :
- distance between centers of adjacent tracer grid cells in the 'y' direction
- units :
- m
- coordinate :
- YG XC
- coverage_content_type :
- modelResult
- comment :
- Alternatively, the length of 'east' side of vorticity grid cells. Note: 'east' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[ 11563.718 , 11593.785 , 11621.304 , ..., 3803.9153, 3573.0938, 3339.542 ], [ 23472.46 , 23533.473 , 23589.314 , ..., 7733.991 , 7266.5024, 6793.6157], [ 23932.863 , 23995.008 , 24051.885 , ..., 7925.047 , 7451.6045, 6973.0957], ..., [ 57150.88 , 57150.88 , 57150.88 , ..., 57150.88 , 57150.88 , 57150.88 ], [ 58072.91 , 58072.91 , 58072.91 , ..., 58072.91 , 58072.91 , 58072.91 ], [ 58725.49 , 58725.49 , 58725.49 , ..., 58725.49 , 58725.49 , 58725.49 ]], [[ 59201.66 , 59201.66 , 59201.66 , ..., 59201.66 , 59201.66 , 59201.66 ], [ 59681.477 , 59681.477 , 59681.477 , ..., 59681.477 , 59681.477 , 59681.477 ], [ 60340.285 , 60340.285 , 60340.285 , ..., 60340.285 , 60340.285 , 60340.285 ], ... [101566.42 , 101566.42 , 101566.42 , ..., 101566.42 , 101566.42 , 101566.42 ], [105206.3 , 105206.3 , 105206.3 , ..., 105206.3 , 105206.3 , 105206.3 ], [107413.53 , 107413.53 , 107413.53 , ..., 107413.53 , 107413.53 , 107413.53 ]], [[108536.33 , 108536.33 , 108536.33 , ..., 108536.33 , 108536.33 , 108536.33 ], [108947.625 , 108947.625 , 108947.625 , ..., 108947.625 , 108947.625 , 108947.625 ], [108940.72 , 108940.72 , 108940.72 , ..., 108940.72 , 108940.72 , 108940.72 ], ..., [ 22091.904 , 23051.049 , 24520.809 , ..., 24520.809 , 23051.049 , 22091.904 ], [ 19147.52 , 20797.248 , 22892.352 , ..., 22892.352 , 20797.248 , 19147.52 ], [ 15461.596 , 18723.23 , 21606.105 , ..., 21606.105 , 18723.23 , 15461.596 ]]], dtype=float32) - rAw(tile, j, i_g)float323.617e+08 3.628e+08 ... 2.922e+08
- long_name :
- area of 'v' grid cell
- units :
- m2
- coordinate :
- YG XC
- coverage_content_type :
- modelResult
- standard_name :
- cell_area
- comment :
- Model 'v' grid cells are staggered in space between adjacent tracer grid cells in the 'x' direction. 'v' grid cell (i,j) is situated at the 'west' edge of tracer grid cell (i, j). Note: 'west' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[3.6169946e+08, 3.6279024e+08, 3.6378880e+08, ..., 6.3726332e+07, 5.6884112e+07, 5.0288944e+07], [3.6904275e+08, 3.7015430e+08, 3.7117190e+08, ..., 6.5421896e+07, 5.8450560e+07, 5.1730956e+07], [3.7659021e+08, 3.7772166e+08, 3.7875747e+08, ..., 6.7598592e+07, 6.0505120e+07, 5.3667840e+07], ..., [3.3636692e+09, 3.3636692e+09, 3.3636692e+09, ..., 3.3636692e+09, 3.3636692e+09, 3.3636692e+09], [3.4631982e+09, 3.4631982e+09, 3.4631982e+09, ..., 3.4631982e+09, 3.4631982e+09, 3.4631982e+09], [3.5442668e+09, 3.5442668e+09, 3.5442668e+09, ..., 3.5442668e+09, 3.5442668e+09, 3.5442668e+09]], [[3.6245120e+09, 3.6245120e+09, 3.6245120e+09, ..., 3.6245120e+09, 3.6245120e+09, 3.6245120e+09], [3.7078205e+09, 3.7078205e+09, 3.7078205e+09, ..., 3.7078205e+09, 3.7078205e+09, 3.7078205e+09], [3.8142915e+09, 3.8142915e+09, 3.8142915e+09, ..., 3.8142915e+09, 3.8142915e+09, 3.8142915e+09], ... [1.1415347e+10, 1.1415347e+10, 1.1415347e+10, ..., 1.1415347e+10, 1.1415347e+10, 1.1415347e+10], [1.1706299e+10, 1.1706299e+10, 1.1706299e+10, ..., 1.1706299e+10, 1.1706299e+10, 1.1706299e+10], [1.1852359e+10, 1.1852359e+10, 1.1852359e+10, ..., 1.1852359e+10, 1.1852359e+10, 1.1852359e+10]], [[1.1896091e+10, 1.1896091e+10, 1.1896091e+10, ..., 1.1896091e+10, 1.1896091e+10, 1.1896091e+10], [1.1875574e+10, 1.1875574e+10, 1.1875574e+10, ..., 1.1875574e+10, 1.1875574e+10, 1.1875574e+10], [1.1817668e+10, 1.1817668e+10, 1.1817668e+10, ..., 1.1817668e+10, 1.1817668e+10, 1.1817668e+10], ..., [4.7981014e+08, 5.1110163e+08, 5.8754176e+08, ..., 6.8245645e+08, 5.8754176e+08, 5.1110163e+08], [3.3588122e+08, 3.9047174e+08, 5.0079645e+08, ..., 6.1789274e+08, 5.0079645e+08, 3.9047174e+08], [1.7369571e+08, 2.9218582e+08, 4.4477142e+08, ..., 5.7864640e+08, 4.4477142e+08, 2.9218582e+08]]], dtype=float32) - rAs(tile, j_g, i)float321.802e+08 1.807e+08 ... 2.8e+08
- long_name :
- area of 'u' grid cell
- units :
- m2
- coverage_content_type :
- modelResult
- standard_name :
- cell_area
- comment :
- Model 'u' grid cells are staggered in space between adjacent tracer grid cells in the 'y' direction. 'u' grid cell (i,j) is situated at the 'south' edge of tracer grid cell (i, j). Note: 'south' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[1.8022141e+08, 1.8074112e+08, 1.8121526e+08, ..., 2.9962764e+07, 2.6617694e+07, 2.3403408e+07], [3.6590867e+08, 3.6696323e+08, 3.6792531e+08, ..., 6.1033988e+07, 5.4247132e+07, 4.7725612e+07], [3.7336256e+08, 3.7443661e+08, 3.7541648e+08, ..., 6.2900624e+07, 5.5990252e+07, 4.9350072e+07], ..., [3.3124155e+09, 3.3124155e+09, 3.3124155e+09, ..., 3.3124155e+09, 3.3124155e+09, 3.3124155e+09], [3.4155474e+09, 3.4155474e+09, 3.4155474e+09, ..., 3.4155474e+09, 3.4155474e+09, 3.4155474e+09], [3.5045770e+09, 3.5045770e+09, 3.5045770e+09, ..., 3.5045770e+09, 3.5045770e+09, 3.5045770e+09]], [[3.5842450e+09, 3.5842450e+09, 3.5842450e+09, ..., 3.5842450e+09, 3.5842450e+09, 3.5842450e+09], [3.6650668e+09, 3.6650668e+09, 3.6650668e+09, ..., 3.6650668e+09, 3.6650668e+09, 3.6650668e+09], [3.7580419e+09, 3.7580419e+09, 3.7580419e+09, ..., 3.7580419e+09, 3.7580419e+09, 3.7580419e+09], ... [1.1204871e+10, 1.1204871e+10, 1.1204871e+10, ..., 1.1204871e+10, 1.1204871e+10, 1.1204871e+10], [1.1581381e+10, 1.1581381e+10, 1.1581381e+10, ..., 1.1581381e+10, 1.1581381e+10, 1.1581381e+10], [1.1794822e+10, 1.1794822e+10, 1.1794822e+10, ..., 1.1794822e+10, 1.1794822e+10, 1.1794822e+10]], [[1.1884406e+10, 1.1884406e+10, 1.1884406e+10, ..., 1.1884406e+10, 1.1884406e+10, 1.1884406e+10], [1.1891903e+10, 1.1891903e+10, 1.1891903e+10, ..., 1.1891903e+10, 1.1891903e+10, 1.1891903e+10], [1.1850071e+10, 1.1850071e+10, 1.1850071e+10, ..., 1.1850071e+10, 1.1850071e+10, 1.1850071e+10], ..., [5.5342240e+08, 6.0073779e+08, 6.7758662e+08, ..., 6.7758662e+08, 6.0073779e+08, 5.5342240e+08], [4.2057974e+08, 4.9237949e+08, 5.9410266e+08, ..., 5.9410266e+08, 4.9237949e+08, 4.2057974e+08], [2.8002694e+08, 4.0091578e+08, 5.3232387e+08, ..., 5.3232387e+08, 4.0091578e+08, 2.8002694e+08]]], dtype=float32) - drC(k_p1)float325.0 10.0 10.0 ... 422.0 445.0 228.2
- long_name :
- distance between the centers of adjacent tracer grid cells in the 'z' direction
- units :
- m
- coverage_content_type :
- modelResult
- comment :
- The first element corresponds to the distance between the depth of the center of the uppermost model grid cell and the surface.
array([ 5. , 10. , 10. , 10. , 10. , 10. , 10. , 10.005, 10.02 , 10.07 , 10.215, 10.56 , 11.28 , 12.59 , 14.73 , 17.93 , 22.335, 27.975, 34.76 , 42.46 , 50.75 , 59.25 , 67.54 , 75.24 , 82.025, 87.665, 92.07 , 95.27 , 97.415, 98.75 , 99.63 , 100.67 , 102.945, 107.945, 117.08 , 130.96 , 149.015, 169.885, 192.19 , 215.025, 238. , 261. , 284. , 307. , 330. , 353. , 376. , 399. , 422. , 445. , 228.25 ], dtype=float32) - drF(k)float3210.0 10.0 10.0 ... 433.5 456.5
- long_name :
- distance between the upper and lower interfaces of the model grid cell
- units :
- m
- coverage_content_type :
- modelResult
- standard_name :
- cell_thickness
- comment :
- Nominal grid cell thickness. Note: in the z* coordinate system used in ECCO V4, actual tracer grid cell thickness, h, varies through time as h(i,j,k,t)= drF(k) hfacC(i,j,k,t).
array([ 10. , 10. , 10. , 10. , 10. , 10. , 10. , 10.01, 10.03, 10.11, 10.32, 10.8 , 11.76, 13.42, 16.04, 19.82, 24.85, 31.1 , 38.42, 46.5 , 55. , 63.5 , 71.58, 78.9 , 85.15, 90.18, 93.96, 96.58, 98.25, 99.25, 100.01, 101.33, 104.56, 111.33, 122.83, 139.09, 158.94, 180.83, 203.55, 226.5 , 249.5 , 272.5 , 295.5 , 318.5 , 341.5 , 364.5 , 387.5 , 410.5 , 433.5 , 456.5 ], dtype=float32) - PHrefC(k)float3249.05 147.1 ... 5.357e+04 5.794e+04
- long_name :
- reference ocean hydrostatic pressure at tracer grid cell center
- units :
- m2 s-2
- coverage_content_type :
- modelResult
- comment :
- PHrefC = p_ref (k) / rhoConst = rhoConst g z(k) / rhoConst = g z(k), where p_ref(k) is reference hydrostatic ocean pressure at center of tracer grid cell k, rhoConst is reference density (1029 kg m-3), g is acceleration due to gravity (9.81 m s-2), and z(k) is depth at center of tracer grid cell k. Units: p:[kg m-1 s-2], rhoConst:[kg m-3], g:[m s-2], z_m(t):[m]. Note: does not include atmospheric pressure loading. Quantity referred to in some contexts as hydrostatic pressure potential. PHIHYDcR is anomaly of PHrefC.
array([4.9049999e+01, 1.4714999e+02, 2.4525000e+02, 3.4335001e+02, 4.4145001e+02, 5.3954999e+02, 6.3765002e+02, 7.3579907e+02, 8.3409528e+02, 9.3288196e+02, 1.0330911e+03, 1.1366847e+03, 1.2473416e+03, 1.3708494e+03, 1.5153507e+03, 1.6912440e+03, 1.9103503e+03, 2.1847852e+03, 2.5257808e+03, 2.9423132e+03, 3.4401709e+03, 4.0214133e+03, 4.6839805e+03, 5.4220850e+03, 6.2267505e+03, 7.0867441e+03, 7.9899507e+03, 8.9245498e+03, 9.8801904e+03, 1.0848928e+04, 1.1826299e+04, 1.2813871e+04, 1.3823762e+04, 1.4882702e+04, 1.6031257e+04, 1.7315975e+04, 1.8777811e+04, 2.0444383e+04, 2.2329768e+04, 2.4439162e+04, 2.6773943e+04, 2.9334352e+04, 3.2120393e+04, 3.5132062e+04, 3.8369363e+04, 4.1832293e+04, 4.5520852e+04, 4.9435043e+04, 5.3574863e+04, 5.7940312e+04], dtype=float32) - PHrefF(k_p1)float320.0 98.1 ... 5.57e+04 6.018e+04
- long_name :
- reference ocean hydrostatic pressure at tracer grid cell top/bottom interface
- units :
- m2 s-2
- coverage_content_type :
- modelResult
- comment :
- PHrefF = p_ref (k_l) / rhoConst = rhoConst g z(k_l) / rhoConst = g z(k_l), where p_ref(k_l) is reference hydrostatic ocean pressure at lower interface of tracer grid cell k, rhoConst is reference density (1029 kg m-3), g is acceleration due to gravity (9.81 m s-2), and z(k) is depth at center of tracer grid cell k. Units: p:[kg m-1 s-2], rhoConst:[kg m-3], g:[m s-2], z_m(t):[m]. Note: does not include atmospheric pressure loading. Quantity referred to in some contexts as hydrostatic pressure potential. See PHrefC
array([ 0. , 98.1 , 196.2 , 294.3 , 392.4 , 490.5 , 588.6 , 686.7 , 784.8981, 883.2924, 982.4715, 1083.7107, 1189.6587, 1305.0243, 1436.6746, 1594.0269, 1788.461 , 2032.2396, 2337.3306, 2714.2307, 3170.3958, 3709.9458, 4332.881 , 5035.0806, 5809.09 , 6644.411 , 7529.0767, 8450.824 , 9398.274 , 10362.106 , 11335.749 , 12316.848 , 13310.895 , 14336.628 , 15428.775 , 16633.738 , 17998.21 , 19557.412 , 21331.355 , 23328.18 , 25550.145 , 27997.74 , 30670.965 , 33569.82 , 36694.305 , 40044.42 , 43620.164 , 47421.54 , 51448.547 , 55701.18 , 60179.445 ], dtype=float32) - hFacC(k, tile, j, i)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- long_name :
- vertical open fraction of tracer grid cell
- coverage_content_type :
- modelResult
- units :
- 1
- comment :
- Tracer grid cells may be fractionally closed in the vertical. The open vertical fraction is hFacC. The model allows for partially-filled cells to represent topographic variations more smoothly (hFacC < 1). Completely closed (dry) tracer grid cells have hFacC = 0. Note: the model z* coordinate system allows hFacC to vary through time. A time-invariant hFacC field is provided for reference.
array([[[[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - hFacW(k, tile, j, i_g)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- long_name :
- vertical open fraction of tracer grid cell 'west' face
- coverage_content_type :
- modelResult
- units :
- 1
- comment :
- The 'west' face of tracer grid cells may be fractionally closed in the vertical. The open vertical fraction is hFacW. The model allows for partially-filled cells for smoother representation of seafloor topography. Tracer grid cells adjacent in the 'x' direction that are partially closed in the vertical have hFacW < 1. The model z* coordinate system used by the model permits hFacC, and therefore hFacW, to vary through time. A time-invariant hFacW field is provided for reference. Note: The term 'west' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - hFacS(k, tile, j_g, i)float320.0 0.0 0.0 0.0 ... 0.0 0.0 0.0 0.0
- long_name :
- vertical open fraction of tracer grid cell 'south' face
- coverage_content_type :
- modelResult
- units :
- 1
- comment :
- The 'south' face of tracer grid cells may be fractionally closed in the vertical. The open vertical fraction is hFacS. The model allows for partially-filled cells for smoother representation of seafloor topography. Tracer grid cells adjacent in the 'y' direction that are partially closed in the vertical have hFacS < 1. The model z* coordinate system used by the model permits hFacC, and therefore hFacS, to vary through time. A time-invariant hFacS field is provided for reference. Note: The term 'south' does not correspond to geographic orientation but is used for convenience to describe the computational grid. See MITgcm documentation for details.
array([[[[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ]], [[1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], [1. , 1. , 1. , ..., 1. , 1. , 1. ], ... [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]], [[0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], ..., [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ], [0. , 0. , 0. , ..., 0. , 0. , 0. ]]]], dtype=float32) - maskC(k, tile, j, i)boolFalse False False ... False False
- long_name :
- wet/dry boolean mask for tracer grid cell
- coverage_content_type :
- modelResult
- comment :
- True for tracer grid cells with nonzero open vertical fraction (hFacC > 0), otherwise False. Although hFacC can vary though time, cells will never close if starting open and will never open if starting closed: hFacC(i,j,k,t) > 0 for all t, if hFacC(i,j,k,t=0) and hFacC(i,j,k,t) = 0 for all t, if hFacC(i,j,k,t=0) = 0. Therefore, maskC is time invariant.
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, False, True]], [[ True, True, True, ..., False, False, True], [ True, True, True, ..., True, False, True], [ True, True, True, ..., True, True, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]]) - maskW(k, tile, j, i_g)boolFalse False False ... False False
- long_name :
- wet/dry boolean mask for 'west' face of tracer grid cell
- coverage_content_type :
- modelResult
- comment :
- True for grid cells with nonzero open vertical fraction along their 'west' face (hFacW > 0), otherwise False. Although hFacW can vary though time, cells will never close if starting open and will never open if starting closed: hFacW(i,j,k,t) > 0 for all t, if hFacW(i,j,k,t=0) and hFacW(i,j,k,t) = 0 for all t, if hFacW(i,j,k,t=0) = 0. Therefore, maskW is time invariant. Note:
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, False]], [[ True, True, True, ..., False, False, False], [ True, True, True, ..., True, False, False], [ True, True, True, ..., True, True, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]]) - maskS(k, tile, j_g, i)boolFalse False False ... False False
- long_name :
- wet/dry boolean mask for 'south' face of tracer grid cell
- coverage_content_type :
- modelResult
- comment :
- True for grid cells with nonzero open vertical fraction along their 'south' face (hFacS > 0), otherwise False. Although hFacS can vary though time, cells will never close if starting open and will never open if starting closed: hFacS(i,j,k,t) > 0 for all t, if hFacS(i,j,k,t=0) and hFacS(i,j,k,t) = 0 for all t, if hFacS(i,j,k,t=0) = 0. Therefore, maskS is time invariant. Note:
array([[[[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True]], [[ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], [ True, True, True, ..., True, True, True], ..., [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, True, True], [ True, True, True, ..., False, False, True]], [[ True, True, True, ..., False, False, True], [ True, True, True, ..., False, False, True], [ True, True, True, ..., True, False, True], ..., ... ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]], [[False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], ..., [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False], [False, False, False, ..., False, False, False]]]])
- long_name :
- Dynamic sea surface height anomaly
- units :
- m
- coverage_content_type :
- modelResult
- standard_name :
- sea_surface_height_above_geoid
- comment :
- Dynamic sea surface height anomaly above the geoid, suitable for comparisons with altimetry sea surface height data products that apply the inverse barometer (IB) correction. Note: SSH is calculated by correcting model sea level anomaly ETAN for three effects: a) global mean steric sea level changes related to density changes in the Boussinesq volume-conserving model (Greatbatch correction, see sterGloH), b) the inverted barometer (IB) effect (see SSHIBC) and c) sea level displacement due to sea-ice and snow pressure loading (see sIceLoad). SSH can be compared with the similarly-named SSH variable in previous ECCO products that did not include atmospheric pressure loading (e.g., Version 4 Release 3). Use SSHNOIBC for comparisons with altimetry data products that do NOT apply the IB correction.
- valid_min :
- -1.8805772066116333
- valid_max :
- 1.4207719564437866
Verifying our new NetCDF files¶
To verify that to_netcdf() worked, load them and compare with the originals.
Compare theta_dataset with dataset_1¶
[10]:
# the first test dataset
dataset_1 = xr.open_dataset(new_filename_1)
# release the file handle (not necessary but generally a good idea)
dataset_1.close()
The np.allclose method does element-by-element comparison of variables
[11]:
# loop through the data variables in dataset_1
for key in dataset_1.keys():
print ('checking %s ' % key)
print ('-- identical in dataset_1 and theta_dataset : %s' % \
np.allclose(dataset_1[key], theta_dataset[key], equal_nan=True))
# note: ``equal_nan`` means nan==nan (default nan != nan)
checking THETA
-- identical in dataset_1 and theta_dataset : True
THETA is the same in both datasets.
Compare custom_dataset with dataset_2¶
[12]:
# our custom dataset
dataset_2 = xr.open_dataset(new_filename_2)
dataset_2.close()
print ('finished loading')
finished loading
[13]:
for key in dataset_2.keys():
print ('checking %s ' % key)
print ('-- identical in dataset_2 and custom_dataset : %s'\
% np.allclose(dataset_2[key], custom_dataset[key], equal_nan=True))
checking SSH
-- identical in dataset_2 and custom_dataset : True
checking THETA
-- identical in dataset_2 and custom_dataset : True
checking CS
-- identical in dataset_2 and custom_dataset : True
checking SN
-- identical in dataset_2 and custom_dataset : True
checking rA
-- identical in dataset_2 and custom_dataset : True
checking dxG
-- identical in dataset_2 and custom_dataset : True
checking dyG
-- identical in dataset_2 and custom_dataset : True
checking Depth
-- identical in dataset_2 and custom_dataset : True
checking rAz
-- identical in dataset_2 and custom_dataset : True
checking dxC
-- identical in dataset_2 and custom_dataset : True
checking dyC
-- identical in dataset_2 and custom_dataset : True
checking rAw
-- identical in dataset_2 and custom_dataset : True
checking rAs
-- identical in dataset_2 and custom_dataset : True
checking drC
-- identical in dataset_2 and custom_dataset : True
checking drF
-- identical in dataset_2 and custom_dataset : True
checking PHrefC
-- identical in dataset_2 and custom_dataset : True
checking PHrefF
-- identical in dataset_2 and custom_dataset : True
checking hFacC
-- identical in dataset_2 and custom_dataset : True
checking hFacW
-- identical in dataset_2 and custom_dataset : True
checking hFacS
-- identical in dataset_2 and custom_dataset : True
checking maskC
-- identical in dataset_2 and custom_dataset : False
checking maskW
-- identical in dataset_2 and custom_dataset : False
checking maskS
-- identical in dataset_2 and custom_dataset : False
SSH and THETA are the same in both datasets, as are most of the grid parameters. But what’s happening with the masks (maskC, maskW, maskS)? Let’s check their data type in each of the datasets:
[14]:
print('custom_dataset.maskC.dtype: ' + str(custom_dataset.maskC.dtype))
print('custom_dataset.maskW.dtype: ' + str(custom_dataset.maskW.dtype))
print('custom_dataset.maskS.dtype: ' + str(custom_dataset.maskS.dtype))
print('dataset_2.maskC.dtype: ' + str(dataset_2.maskC.dtype))
print('dataset_2.maskW.dtype: ' + str(dataset_2.maskW.dtype))
print('dataset_2.maskS.dtype: ' + str(dataset_2.maskS.dtype))
custom_dataset.maskC.dtype: bool
custom_dataset.maskW.dtype: bool
custom_dataset.maskS.dtype: bool
dataset_2.maskC.dtype: float32
dataset_2.maskW.dtype: float32
dataset_2.maskS.dtype: float32
So for some reason after these boolean (bool) fields were saved in the file, they were re-opened as float32 floating-point numbers. Let’s fix this and then compare the masks again.
[15]:
dataset_2.maskC.data = dataset_2.maskC.data.astype('bool')
dataset_2.maskW.data = dataset_2.maskW.data.astype('bool')
dataset_2.maskS.data = dataset_2.maskS.data.astype('bool')
for key in dataset_2.keys():
print ('checking %s ' % key)
print ('-- identical in dataset_2 and custom_dataset : %s'\
% np.allclose(dataset_2[key], custom_dataset[key], equal_nan=True))
checking SSH
-- identical in dataset_2 and custom_dataset : True
checking THETA
-- identical in dataset_2 and custom_dataset : True
checking CS
-- identical in dataset_2 and custom_dataset : True
checking SN
-- identical in dataset_2 and custom_dataset : True
checking rA
-- identical in dataset_2 and custom_dataset : True
checking dxG
-- identical in dataset_2 and custom_dataset : True
checking dyG
-- identical in dataset_2 and custom_dataset : True
checking Depth
-- identical in dataset_2 and custom_dataset : True
checking rAz
-- identical in dataset_2 and custom_dataset : True
checking dxC
-- identical in dataset_2 and custom_dataset : True
checking dyC
-- identical in dataset_2 and custom_dataset : True
checking rAw
-- identical in dataset_2 and custom_dataset : True
checking rAs
-- identical in dataset_2 and custom_dataset : True
checking drC
-- identical in dataset_2 and custom_dataset : True
checking drF
-- identical in dataset_2 and custom_dataset : True
checking PHrefC
-- identical in dataset_2 and custom_dataset : True
checking PHrefF
-- identical in dataset_2 and custom_dataset : True
checking hFacC
-- identical in dataset_2 and custom_dataset : True
checking hFacW
-- identical in dataset_2 and custom_dataset : True
checking hFacS
-- identical in dataset_2 and custom_dataset : True
checking maskC
-- identical in dataset_2 and custom_dataset : True
checking maskW
-- identical in dataset_2 and custom_dataset : True
checking maskS
-- identical in dataset_2 and custom_dataset : True
Now we can confirm that all the variables in the datasets match!
Saving the results of calculations¶
Calculations in the form of DataArrays¶
Often we would like to store the results of our calculations to disk. If your operations are made at the level of DataArray objects (and not the lower ndarray level) then you can use these same methods to save your results. All of the coordinates will be preserved (although attributes be lost). Let’s demonstrate by making a dummy calculation on SSH
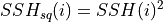
[16]:
SSH_sq = custom_dataset.SSH * custom_dataset.SSH
SSH_sq
[16]:
<xarray.DataArray 'SSH' (time: 1, tile: 3, j: 90, i: 90)>
array([[[[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[2.0671911 , 2.0848224 , 2.0841086 , ..., 1.7935177 ,
1.7659562 , 1.739012 ],
[1.9880351 , 2.0051208 , 2.002616 , ..., 1.7589175 ,
1.7311752 , 1.7012385 ],
[1.8994561 , 1.9122815 , 1.9098864 , ..., 1.7312655 ,
1.7034038 , 1.6693714 ]],
[[1.8020678 , 1.810736 , 1.8133253 , ..., 1.7026882 ,
1.6756296 , 1.6379843 ],
[1.7121627 , 1.719325 , 1.7317982 , ..., 1.6670871 ,
1.6423833 , 1.6031213 ],
[1.6535913 , 1.6606174 , 1.6836028 , ..., 1.6204541 ,
1.5996994 , 1.5617636 ],
...
[0.0358197 , 0.03125747, 0.02652959, ..., nan,
0.28220004, 0.31124577],
[0.02917161, 0.02486313, 0.02061234, ..., nan,
0.2443076 , 0.2719567 ],
[0.02321114, 0.01939201, 0.01575851, ..., nan,
nan, 0.23898326]],
[[0.01884312, 0.01536459, 0.01223672, ..., nan,
nan, 0.2193556 ],
[0.01608207, 0.01280527, 0.0099735 , ..., 0.151336 ,
nan, 0.20633157],
[0.01495756, 0.01177618, 0.00917115, ..., 0.16071834,
0.1699125 , 0.20213497],
...,
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan]]]], dtype=float32)
Coordinates:
* i (i) int32 0 1 2 3 4 5 6 7 8 9 10 ... 80 81 82 83 84 85 86 87 88 89
* j (j) int32 0 1 2 3 4 5 6 7 8 9 10 ... 80 81 82 83 84 85 86 87 88 89
* tile (tile) int32 0 1 2
* time (time) datetime64[ns] 2010-03-16T12:00:00
XC (tile, j, i) float32 -111.6 -111.3 -110.9 ... 50.97 51.44 51.84
YC (tile, j, i) float32 -88.24 -88.38 -88.52 ... 67.62 67.53 67.47- time: 1
- tile: 3
- j: 90
- i: 90
- nan nan nan nan nan nan nan nan ... 0.2272 nan nan nan nan nan nan
array([[[[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [2.0671911 , 2.0848224 , 2.0841086 , ..., 1.7935177 , 1.7659562 , 1.739012 ], [1.9880351 , 2.0051208 , 2.002616 , ..., 1.7589175 , 1.7311752 , 1.7012385 ], [1.8994561 , 1.9122815 , 1.9098864 , ..., 1.7312655 , 1.7034038 , 1.6693714 ]], [[1.8020678 , 1.810736 , 1.8133253 , ..., 1.7026882 , 1.6756296 , 1.6379843 ], [1.7121627 , 1.719325 , 1.7317982 , ..., 1.6670871 , 1.6423833 , 1.6031213 ], [1.6535913 , 1.6606174 , 1.6836028 , ..., 1.6204541 , 1.5996994 , 1.5617636 ], ... [0.0358197 , 0.03125747, 0.02652959, ..., nan, 0.28220004, 0.31124577], [0.02917161, 0.02486313, 0.02061234, ..., nan, 0.2443076 , 0.2719567 ], [0.02321114, 0.01939201, 0.01575851, ..., nan, nan, 0.23898326]], [[0.01884312, 0.01536459, 0.01223672, ..., nan, nan, 0.2193556 ], [0.01608207, 0.01280527, 0.0099735 , ..., 0.151336 , nan, 0.20633157], [0.01495756, 0.01177618, 0.00917115, ..., 0.16071834, 0.1699125 , 0.20213497], ..., [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan]]]], dtype=float32) - i(i)int320 1 2 3 4 5 6 ... 84 85 86 87 88 89
- axis :
- X
- long_name :
- grid index in x for variables at tracer and 'v' locations
- swap_dim :
- XC
- comment :
- In the Arakawa C-grid system, tracer (e.g., THETA) and 'v' variables (e.g., VVEL) have the same x coordinate on the model grid.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j(j)int320 1 2 3 4 5 6 ... 84 85 86 87 88 89
- axis :
- Y
- long_name :
- grid index in y for variables at tracer and 'u' locations
- swap_dim :
- YC
- comment :
- In the Arakawa C-grid system, tracer (e.g., THETA) and 'u' variables (e.g., UVEL) have the same y coordinate on the model grid.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - tile(tile)int320 1 2
- long_name :
- lat-lon-cap tile index
- comment :
- The ECCO V4 horizontal model grid is divided into 13 tiles of 90x90 cells for convenience.
- coverage_content_type :
- coordinate
array([0, 1, 2])
- time(time)datetime64[ns]2010-03-16T12:00:00
- long_name :
- center time of averaging period
- axis :
- T
- bounds :
- time_bnds
- coverage_content_type :
- coordinate
- standard_name :
- time
array(['2010-03-16T12:00:00.000000000'], dtype='datetime64[ns]')
- XC(tile, j, i)float32-111.6 -111.3 ... 51.44 51.84
- long_name :
- longitude of tracer grid cell center
- units :
- degrees_east
- coordinate :
- YC XC
- bounds :
- XC_bnds
- comment :
- nonuniform grid spacing
- coverage_content_type :
- coordinate
- standard_name :
- longitude
array([[[-111.60647 , -111.303 , -110.94285 , ..., 64.791115, 64.80521 , 64.81917 ], [-104.8196 , -103.928444, -102.87706 , ..., 64.36745 , 64.41012 , 64.4524 ], [ -98.198784, -96.788055, -95.14185 , ..., 63.936497, 64.008224, 64.0793 ], ..., [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ]], [[ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], ... [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ]], [[ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], ..., [ -37.730072, -37.17829 , -36.597565, ..., 50.597565, 51.17829 , 51.730072], [ -37.771988, -37.291943, -36.764027, ..., 50.764027, 51.291943, 51.771988], [ -37.837925, -37.44421 , -36.968143, ..., 50.968143, 51.44421 , 51.837925]]], dtype=float32) - YC(tile, j, i)float32-88.24 -88.38 ... 67.53 67.47
- long_name :
- latitude of tracer grid cell center
- units :
- degrees_north
- coordinate :
- YC XC
- bounds :
- YC_bnds
- comment :
- nonuniform grid spacing
- coverage_content_type :
- coordinate
- standard_name :
- latitude
array([[[-88.24259 , -88.382515 , -88.52242 , ..., -80.57383 , -80.50492 , -80.43992 ], [-88.21676 , -88.354485 , -88.49178 , ..., -80.56925 , -80.50038 , -80.43542 ], [-88.164764 , -88.29827 , -88.43065 , ..., -80.55984 , -80.49105 , -80.42617 ], ..., [-58.321964 , -58.321964 , -58.321964 , ..., -58.321964 , -58.321964 , -58.321964 ], [-57.79962 , -57.79962 , -57.79962 , ..., -57.79962 , -57.79962 , -57.79962 ], [-57.271408 , -57.271408 , -57.271408 , ..., -57.271408 , -57.271408 , -57.271408 ]], [[-56.73891 , -56.73891 , -56.73891 , ..., -56.73891 , -56.73891 , -56.73891 ], [-56.2021 , -56.2021 , -56.2021 , ..., -56.2021 , -56.2021 , -56.2021 ], [-55.65936 , -55.65936 , -55.65936 , ..., -55.65936 , -55.65936 , -55.65936 ], ... [ 7.5699615, 7.5699615, 7.5699615, ..., 7.5699615, 7.5699615, 7.5699615], [ 8.516253 , 8.516253 , 8.516253 , ..., 8.516253 , 8.516253 , 8.516253 ], [ 9.482398 , 9.482398 , 9.482398 , ..., 9.482398 , 9.482398 , 9.482398 ]], [[ 10.458642 , 10.458642 , 10.458642 , ..., 10.458642 , 10.458642 , 10.458642 ], [ 11.438585 , 11.438585 , 11.438585 , ..., 11.438585 , 11.438585 , 11.438585 ], [ 12.418467 , 12.418467 , 12.418467 , ..., 12.418467 , 12.418467 , 12.418467 ], ..., [ 67.16407 , 67.19428 , 67.24966 , ..., 67.24966 , 67.19428 , 67.16407 ], [ 67.33552 , 67.37607 , 67.445274 , ..., 67.445274 , 67.37607 , 67.33552 ], [ 67.47211 , 67.53387 , 67.62317 , ..., 67.62317 , 67.53387 , 67.47211 ]]], dtype=float32)
SSH_sq is itself a DataArray.
Before saving, let’s give our new SSH_sq variable a better name and descriptive attributes.
[17]:
SSH_sq.name = 'SSH^2'
SSH_sq.attrs['long_name'] = 'Square of Surface Height Anomaly'
SSH_sq.attrs['units'] = 'm^2'
# Let's see the result
SSH_sq
[17]:
<xarray.DataArray 'SSH^2' (time: 1, tile: 3, j: 90, i: 90)>
array([[[[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[2.0671911 , 2.0848224 , 2.0841086 , ..., 1.7935177 ,
1.7659562 , 1.739012 ],
[1.9880351 , 2.0051208 , 2.002616 , ..., 1.7589175 ,
1.7311752 , 1.7012385 ],
[1.8994561 , 1.9122815 , 1.9098864 , ..., 1.7312655 ,
1.7034038 , 1.6693714 ]],
[[1.8020678 , 1.810736 , 1.8133253 , ..., 1.7026882 ,
1.6756296 , 1.6379843 ],
[1.7121627 , 1.719325 , 1.7317982 , ..., 1.6670871 ,
1.6423833 , 1.6031213 ],
[1.6535913 , 1.6606174 , 1.6836028 , ..., 1.6204541 ,
1.5996994 , 1.5617636 ],
...
[0.0358197 , 0.03125747, 0.02652959, ..., nan,
0.28220004, 0.31124577],
[0.02917161, 0.02486313, 0.02061234, ..., nan,
0.2443076 , 0.2719567 ],
[0.02321114, 0.01939201, 0.01575851, ..., nan,
nan, 0.23898326]],
[[0.01884312, 0.01536459, 0.01223672, ..., nan,
nan, 0.2193556 ],
[0.01608207, 0.01280527, 0.0099735 , ..., 0.151336 ,
nan, 0.20633157],
[0.01495756, 0.01177618, 0.00917115, ..., 0.16071834,
0.1699125 , 0.20213497],
...,
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan]]]], dtype=float32)
Coordinates:
* i (i) int32 0 1 2 3 4 5 6 7 8 9 10 ... 80 81 82 83 84 85 86 87 88 89
* j (j) int32 0 1 2 3 4 5 6 7 8 9 10 ... 80 81 82 83 84 85 86 87 88 89
* tile (tile) int32 0 1 2
* time (time) datetime64[ns] 2010-03-16T12:00:00
XC (tile, j, i) float32 -111.6 -111.3 -110.9 ... 50.97 51.44 51.84
YC (tile, j, i) float32 -88.24 -88.38 -88.52 ... 67.62 67.53 67.47
Attributes:
long_name: Square of Surface Height Anomaly
units: m^2- time: 1
- tile: 3
- j: 90
- i: 90
- nan nan nan nan nan nan nan nan ... 0.2272 nan nan nan nan nan nan
array([[[[ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], ..., [2.0671911 , 2.0848224 , 2.0841086 , ..., 1.7935177 , 1.7659562 , 1.739012 ], [1.9880351 , 2.0051208 , 2.002616 , ..., 1.7589175 , 1.7311752 , 1.7012385 ], [1.8994561 , 1.9122815 , 1.9098864 , ..., 1.7312655 , 1.7034038 , 1.6693714 ]], [[1.8020678 , 1.810736 , 1.8133253 , ..., 1.7026882 , 1.6756296 , 1.6379843 ], [1.7121627 , 1.719325 , 1.7317982 , ..., 1.6670871 , 1.6423833 , 1.6031213 ], [1.6535913 , 1.6606174 , 1.6836028 , ..., 1.6204541 , 1.5996994 , 1.5617636 ], ... [0.0358197 , 0.03125747, 0.02652959, ..., nan, 0.28220004, 0.31124577], [0.02917161, 0.02486313, 0.02061234, ..., nan, 0.2443076 , 0.2719567 ], [0.02321114, 0.01939201, 0.01575851, ..., nan, nan, 0.23898326]], [[0.01884312, 0.01536459, 0.01223672, ..., nan, nan, 0.2193556 ], [0.01608207, 0.01280527, 0.0099735 , ..., 0.151336 , nan, 0.20633157], [0.01495756, 0.01177618, 0.00917115, ..., 0.16071834, 0.1699125 , 0.20213497], ..., [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan], [ nan, nan, nan, ..., nan, nan, nan]]]], dtype=float32) - i(i)int320 1 2 3 4 5 6 ... 84 85 86 87 88 89
- axis :
- X
- long_name :
- grid index in x for variables at tracer and 'v' locations
- swap_dim :
- XC
- comment :
- In the Arakawa C-grid system, tracer (e.g., THETA) and 'v' variables (e.g., VVEL) have the same x coordinate on the model grid.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - j(j)int320 1 2 3 4 5 6 ... 84 85 86 87 88 89
- axis :
- Y
- long_name :
- grid index in y for variables at tracer and 'u' locations
- swap_dim :
- YC
- comment :
- In the Arakawa C-grid system, tracer (e.g., THETA) and 'u' variables (e.g., UVEL) have the same y coordinate on the model grid.
- coverage_content_type :
- coordinate
array([ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89]) - tile(tile)int320 1 2
- long_name :
- lat-lon-cap tile index
- comment :
- The ECCO V4 horizontal model grid is divided into 13 tiles of 90x90 cells for convenience.
- coverage_content_type :
- coordinate
array([0, 1, 2])
- time(time)datetime64[ns]2010-03-16T12:00:00
- long_name :
- center time of averaging period
- axis :
- T
- bounds :
- time_bnds
- coverage_content_type :
- coordinate
- standard_name :
- time
array(['2010-03-16T12:00:00.000000000'], dtype='datetime64[ns]')
- XC(tile, j, i)float32-111.6 -111.3 ... 51.44 51.84
- long_name :
- longitude of tracer grid cell center
- units :
- degrees_east
- coordinate :
- YC XC
- bounds :
- XC_bnds
- comment :
- nonuniform grid spacing
- coverage_content_type :
- coordinate
- standard_name :
- longitude
array([[[-111.60647 , -111.303 , -110.94285 , ..., 64.791115, 64.80521 , 64.81917 ], [-104.8196 , -103.928444, -102.87706 , ..., 64.36745 , 64.41012 , 64.4524 ], [ -98.198784, -96.788055, -95.14185 , ..., 63.936497, 64.008224, 64.0793 ], ..., [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ]], [[ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], ... [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ]], [[ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], [ -37.5 , -36.5 , -35.5 , ..., 49.5 , 50.5 , 51.5 ], ..., [ -37.730072, -37.17829 , -36.597565, ..., 50.597565, 51.17829 , 51.730072], [ -37.771988, -37.291943, -36.764027, ..., 50.764027, 51.291943, 51.771988], [ -37.837925, -37.44421 , -36.968143, ..., 50.968143, 51.44421 , 51.837925]]], dtype=float32) - YC(tile, j, i)float32-88.24 -88.38 ... 67.53 67.47
- long_name :
- latitude of tracer grid cell center
- units :
- degrees_north
- coordinate :
- YC XC
- bounds :
- YC_bnds
- comment :
- nonuniform grid spacing
- coverage_content_type :
- coordinate
- standard_name :
- latitude
array([[[-88.24259 , -88.382515 , -88.52242 , ..., -80.57383 , -80.50492 , -80.43992 ], [-88.21676 , -88.354485 , -88.49178 , ..., -80.56925 , -80.50038 , -80.43542 ], [-88.164764 , -88.29827 , -88.43065 , ..., -80.55984 , -80.49105 , -80.42617 ], ..., [-58.321964 , -58.321964 , -58.321964 , ..., -58.321964 , -58.321964 , -58.321964 ], [-57.79962 , -57.79962 , -57.79962 , ..., -57.79962 , -57.79962 , -57.79962 ], [-57.271408 , -57.271408 , -57.271408 , ..., -57.271408 , -57.271408 , -57.271408 ]], [[-56.73891 , -56.73891 , -56.73891 , ..., -56.73891 , -56.73891 , -56.73891 ], [-56.2021 , -56.2021 , -56.2021 , ..., -56.2021 , -56.2021 , -56.2021 ], [-55.65936 , -55.65936 , -55.65936 , ..., -55.65936 , -55.65936 , -55.65936 ], ... [ 7.5699615, 7.5699615, 7.5699615, ..., 7.5699615, 7.5699615, 7.5699615], [ 8.516253 , 8.516253 , 8.516253 , ..., 8.516253 , 8.516253 , 8.516253 ], [ 9.482398 , 9.482398 , 9.482398 , ..., 9.482398 , 9.482398 , 9.482398 ]], [[ 10.458642 , 10.458642 , 10.458642 , ..., 10.458642 , 10.458642 , 10.458642 ], [ 11.438585 , 11.438585 , 11.438585 , ..., 11.438585 , 11.438585 , 11.438585 ], [ 12.418467 , 12.418467 , 12.418467 , ..., 12.418467 , 12.418467 , 12.418467 ], ..., [ 67.16407 , 67.19428 , 67.24966 , ..., 67.24966 , 67.19428 , 67.16407 ], [ 67.33552 , 67.37607 , 67.445274 , ..., 67.445274 , 67.37607 , 67.33552 ], [ 67.47211 , 67.53387 , 67.62317 , ..., 67.62317 , 67.53387 , 67.47211 ]]], dtype=float32)
- long_name :
- Square of Surface Height Anomaly
- units :
- m^2
much better! Now we’ll save.
[18]:
new_filename_3 = './ssh_sq_DataArray.nc'
print ('saving to ', new_filename_3)
SSH_sq.to_netcdf(path=new_filename_3)
print ('finished saving')
saving to ./ssh_sq_DataArray.nc
finished saving
Calculations in the form of numpy ndarrays¶
If calculations are made at the ndarray level then the results will also be ndarrays.
[19]:
SSH_dummy_ndarray = custom_dataset.SSH.values * custom_dataset.SSH.values
type(SSH_dummy_ndarray)
[19]:
numpy.ndarray
You’ll need to use different methods to save these results to NetCDF files, one of which is described here: http://pyhogs.github.io/intro_netcdf4.html
Summary¶
Saving Datasets and DataArrays to disk as NetCDF files is fun and easy with xarray!